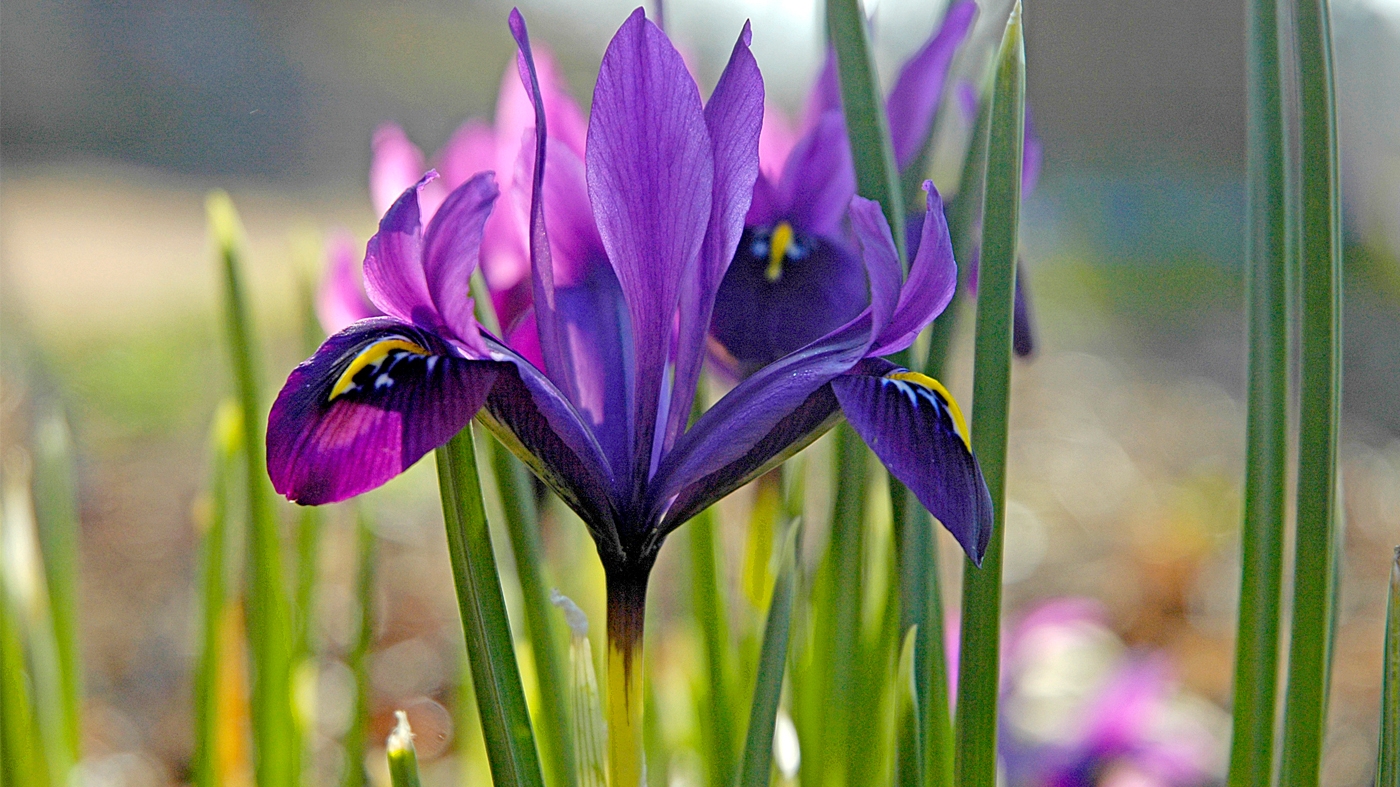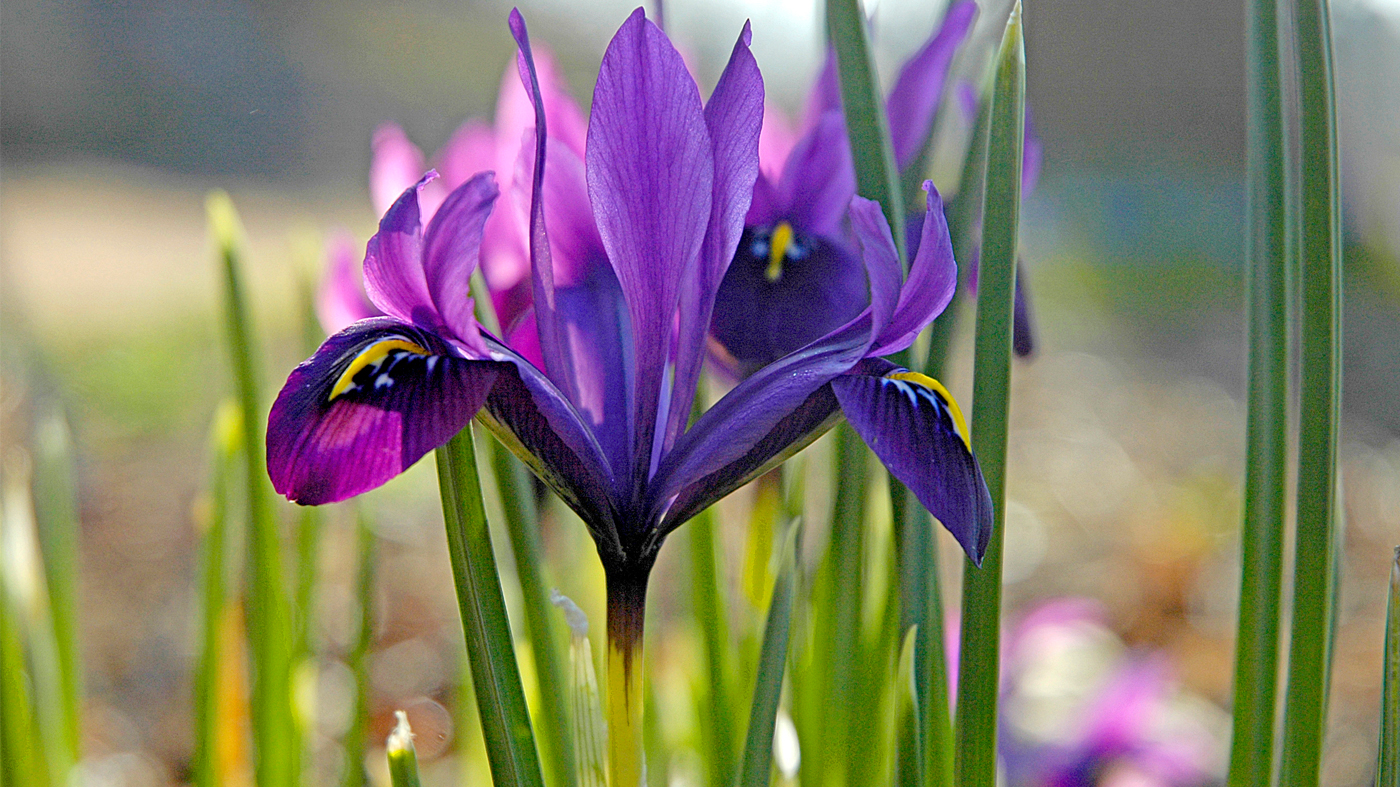

Wildlife
Garden Stories
Studying the cryptic and beautiful Mompha moths
Most butterflies and moths featured in popular magazines and other media are large, well-known species, such as monarchs and luna moths.
Within scientific communities as well, species descriptions are biased toward larger moths, overlooking the multitude of tiny ones. Despite this tendency to favor larger species, the average moth is actually quite small, though far from nondescript!

Mompha species moth; photo taken in Utah
My research at the Chicago Botanic Garden focuses on an insufficiently studied moth group called Mompha, the largest genus within the family Momphidae. Mompha are tiny moths characterized by 4- to 8-millimeter tufted forewings and distinct color patterns.
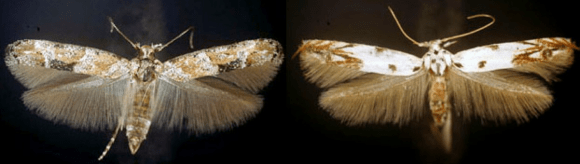
Specimens up close: Mompha stellella on the left and Mompha eloisella on the right. Both are found in Illinois, typically during the month of August. Photo credit: Terry Harrison
In North America, there are approximately 40 described species, or taxa, of Mompha. In addition to these identified species, a number of undescribed taxa are located throughout the North American West and Southwest. Mompha larvae feed on the reproductive (i.e., flowers, buds, and fruits) and vegetative (i.e., leaves, stems, and roots) structures of members of the Lythraceae, Cistaceae, Rubiaceae, and, most commonly, Onagraceae (evening primroses). In Illinois, Mompha can be collected in your backyard from Oenothera biennis (common evening primrose).

Examples of Mompha bud-feeding and Mompha fruit-feeding caterpillars
Because many Mompha species share the same coloration, the only morphological characteristics—size, shape, and structure of an organism or one of its parts—that accurately differentiate taxa are unique genitalia. Experienced lepidopterists—butterfly and moth researchers or collectors—are able to carefully dissect moths in order to view their genitalia. However, due to the unique skills involved in moth dissection and genitalia identification, few scientists are qualified to identify different Mompha species.

Close-up of Mompha species caterpillar
Instead of conducting genitalia dissections, I am sequencing six genes from hundreds of Mompha collected over the span of three years from the Western and Southwestern United States. DNA, like morphological characteristics, can be used to identify and characterize differences between species. To analyze the differences within Mompha DNA, I modeled phylogenetic trees.
Phylogenetic trees depict evolutionary relationships between species in regard to genetic characteristic; closely related species share similar DNA and are thus placed close together on a phylogenetic tree. These trees will allow me to describe the natural history of Mompha in North America. This means that I will be able to identify new Mompha species, as well as Mompha host plant preferences, plant structure preferences, emergence times, and geographic isolation.

